
We use cookies to ensure our website works properly and to personalise your experience. Cookies policy

We use cookies to ensure our website works properly and to personalise your experience. Cookies policy
Krantiveer Vasantrao Narayanrao Naik Shikshan Prasarak Sanstha's, Institute of Pharmaceutical Education & Research, Canada Corner, Nashik - (422002) Maharashtra, India
Drug development for the treatment of disease is being revolutionised by RNA-based oligonucleotide therapies. This class of drugs differs from small molecule and protein therapies in several ways, such as how they work and how they are related to clinical pharmacology. Since the COVID-19 mRNA vaccine was approved and nucleoside base alterations won the Nobel Prize in 2023, RNA treatments have gained attention and are revolutionising the drug development process. Although the phrase "RNA therapeutics" has been applied in several settings, the focus of this review is on therapies that target RNA or employ RNA as a component of RNA to achieve therapeutic effects. Small interfering RNA (siRNA) treatments have gained popularity across a variety of therapeutic areas since the first siRNA treatment was licenced in 2018.Thus, it would be beneficial for many parties going forward to do a detailed review of the clinical pharmacology of siRNA therapies with FDA approval. This review covers not only ADME characteristics but also clinical pharmacology data on RNA treatments, such as population pharmacokinetic studies and drug-drug interactions. With the anticipated high growth of the RNA therapies market, thorough understanding will be essential to interpreting and assessing the pharmacological properties.
RNA treatments have led the way in several medical applications in the last few times, including vaccinations against the SARS-CoV-2 virus and uncommon illnesses involving gene [removed]COVID-19). The unique target specificity of RNA treatments can offer important advantages over traditional pharmaceuticals by altering the translation of disease-causing proteins by downregulating a gene target or by injecting synthetic mRNA to translate the encoded target protein (kole et al.,2012, Aagaard) (rossi et al.,2007). Given that siRNA was first discovered in 19983, the 2018 licencing of the first short interfering RNA (siRNA) medication, patisiran, highlights the rapid therapeutic advancement in a short period of 20 years(Adams et al.,2018).Two COVID-19 mRNA vaccines were approved for emergency use in 2020 and were administered to millions of patients worldwide, marking an important historical milestone as the first widely utilised RNA medications (Baden et al.,2021) (polack et al.,2020)? .On the left is a diagram showing the RNAi pathway and the production of siRNAs. On the right-hand side are various entrance points for RNA- or DNA-based RNAi treatments that penetrate cells and are loaded into RISC to encourage homology-dependent destruction of target mRNA. Viral or non-viral vectors that encode Pol II-driven miRNA mimics or Pol III-driven shRNA can be used to deliver DNA-based therapeutics. RNA interference (RNAi)-based therapeutics can be delivered in a number of forms, including complexed in gold, lipid, or polymer nanoparticles; alternatively, they can be given as synthesised or GalNAc-conjugated dier-substrate siRNA or modified or unmodified siRNA. In mammalian cells, 2 nt 30 overhangs may cause RNAi-based PTGS (Elbashir et al.,2001) (sm et al.,
? ? ? ?
? ? ? ? ? ? 
? ? ? ?
Figure 1. The RNAi mechanism and entry points of RNAi therapeutics (corydon et al.,2023)
In 2004, the first clinical trial employing siRNA-based drug siRNA-027 was initiated for the treatment of age-related macular degeneration (AMD) (kaiser et al.,2010) (corydon et al.,2023a).
? ? ? ?
? ? ? ? ? ? 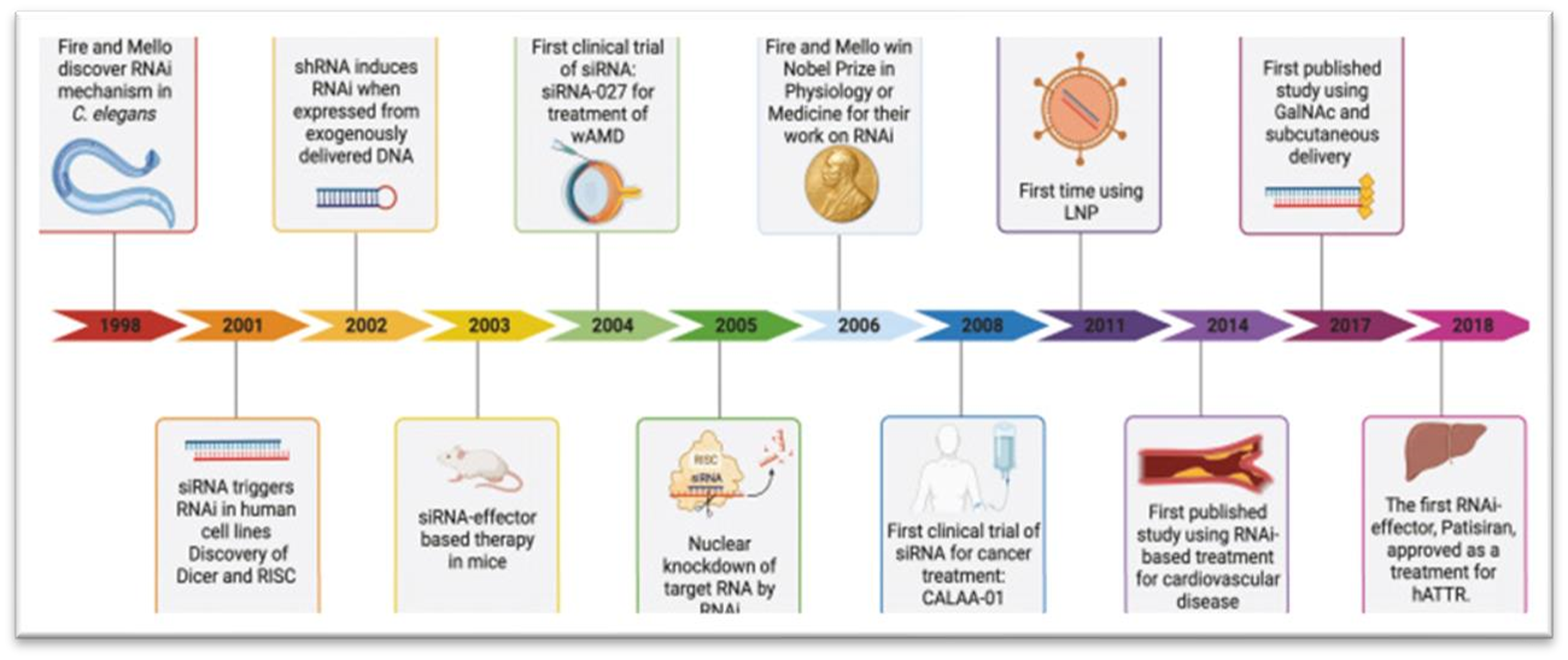
? ? ? ?
Figure 2: Development of RNAi-based: Milestones: therapeutics (corydon et al.,2023b).
CLASSIFICATION
? ? ? ?
? ? ? ? ? ? 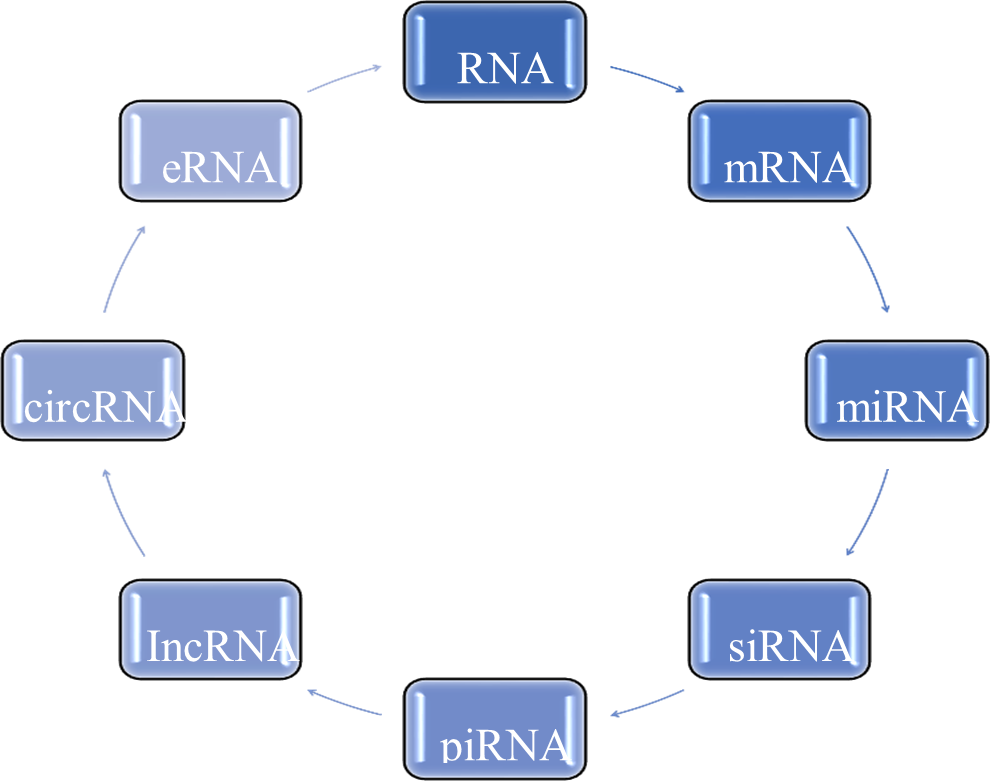
? ? ? ?
Figure 3: Classification of m-RNA
CODING RNA- mRNA [messenger RNA]
Since it was originally discovered that mRNA functions as templates for the translation of proteins, their biological relevance has long been understood. RNA polymerase is normally used to transcribe pre-mRNA (cole et al., 2001).
SMALL CODIND RNA
miRNA and siRNA
Non-coding RNAs (non-coding siRNAs) like miRNA and siRNA are mostly studied for their capacity to control gene expression, mainly via silencing genes. Their length varies between 20 and 35 nucleotides. While miRNAs are endogenous and expressed from an organism's DNA, siRNAs are widely thought to be mostly foreign in origin, originating from viruses, transposons, or transgenic triggers (rw et al.,2009). The degree of complementarity between the sequences determines the regulatory action that follows target binding. Usually, siRNA drives RISC to precisely matched targets and causes target destruction (ryther et al.,2005). miRNA often interacts with the 3' untranslated region (UTR) of mRNA, occasionally causing different bulges and mismatches. miRNA nucleotides 2?8 represent the seed region, which is necessary for target base pairing and identification (rw et al.,2009a).
PiRNA
With a length of 21?35 nucleotides, piRNA is a kind of short non-coding RNA that is exclusive to animals. Its main functions include regulating gene expression, suppressing transposable elements, particularly in the germline, and preventing viral infection (ozata et al.,2019). Piwi clusters are unique genomic loci where piRNA-encoding genes are found.Pol II transcribes these genes to yield more than 90% of the single-stranded primary piRNAs(chen et al.,2021).
LONG NONCODING RNA [IncRNA]
Longer than 200 nucleotides are the normal length of lncRNAs, which set them apart from smaller ncRNAs like tRNA and miRNA. They can be spliced and polyadenylated similarly to mRNA transcripts, albeit it is not required. Their primary transcription is carried out by RNA pol II (mattick et al.,2023).????????????????????????????????????????????????????????????????????????
Circular RNA [circRNA]
A kind of lncRNA known as "circRNA" has a covalently enclosed loop structure that gives it intrinsic resistance to exonuclease degradation and more stability than linear RNA (enuka et al.,2016; Zhang et al.,2016).
Enhancer RNA [eRNA]
Global profiling and annotation indicate that 40,000?65,000 eRNAs are expressed in human cells (Andersson et al.,2014) (arner et al.,2015) (meers et al.,2018) (chen et al.,2013).??????????????????????????????????????????????????????????????????????????????
Absorption and Administration Routes
?Parenterally administered RNA treatments are frequently administered intravenously or subcutaneously. Because of nucleolytic degradation, RNA treatments have low to moderate bioavailability after subcutaneous delivery when compared to other drugs (ovacik et al.,2018) (keizer et al.,2010). For instance, it was shown that inclisiran's systemic bioavailability in rats was 35.1?48.9% and in monkeys, 24.7?33.8%, following subcutaneous treatment(EMA et al.,2022).????????????????????????????????????????????????? ????????????????????????????????????????????????????????????????????????????????????
Distribution
In the cytoplasm or nucleus of cells, RNA therapies interact with their RNA targets (Liberman et al.,2018). As the primary excretion and metabolic organs, the kidney and liver receive a systemic distribution of most RNA therapies(amantana and lversen et al.,2005)(sands et al.,1994)(rosie et al.,2004).
Metabolism:
Exo- and endonucleases are typical mediators of the degradation of RNA therapies. Generally speaking, 3-exonucleases break down unmodified oligonucleotides quickly (EDER et al.,1991) (wojcik et al.,2007) (dirin and winkler et al.,2013). Since the liver is where most drug processing occurs, medicines are mostly transported to this organ out of all the others. Certain RNA therapeutics have been examined using recombinant cytochrome P450 (CYP) systems, liver microsomes, or the s9 part of the liver to better understand how each medicine is metabolised. Clinical interactions with CYP enzymes are uncommon because the majority of GalNAc siRNA medicines are not CYP enzyme substrates (Ramsden et al.,2019).??????????????????????????????????????????????????????????????????????
Elimination
Each RNA therapy has distinct excretion-related pharmacokinetic characteristics, even though renal clearance accounts for the majority of RNA treatments' primary excretion. For example, the half-life of fomivirsen, which is cleared from the vitreous humour at a first-order clearance rate, is around 55 hours. Folimivirsen levels in the vitreous humour may decrease over time due to absorption into the retina and other ocular tissues or nucleases metabolising the compound (geary et al.,2002). In addition, during the first 24 hours after a single intake, mipomersen excretes relatively little in the urine. People specifically excrete 1.38 to 3.30% throughout the course of the first 24 hours following a 2-hour intravenous infusion (bell et al ,. 2011).Research utilising the 5/6 nephrectomy rat model, which symbolises moderate to severe renal impairment, revealed that neither the liver's PK profile nor the outcomes associated with Parkinson's disease were significantly affected by a decrease in urine production(mcdougall et al.,2022).? ?
Pharmacokinetic-Pharmacodynamic PK/PD Relationships
By creating a link between dose-concentration relationships (pharmacokinetics, PK) and concentration-effect relationships (pharmacodynamics, PD), PK/PD analyses calculate the effect of pharmaceutical doses across time (derendorf and meibohm et al.,1999). Although PD markers for RNA therapeutics are well-established, the PK/PD connection is more complex than it is for conventional small drugs. Additionally, biologics display the intricate PK/PD interaction, which was clarified by the target-mediated drug disposal mechanism (an et al.,2020) Alternative PK profiles need to be identified in order to resolve this difference in the plasma profile with PD activity. It was found that PD activity is more closely correlated with drug concentration in organs than plasma concentration in cases where RNA treatments target specific organs (Shimizu et L.,2015). Since RISC is a structural prerequisite for the activity of siRNA medicines, PD effect profiles in nonclinical trials substantially more closely match RISC concentration (mcdougall et al.,2022a).??????????????????????????????????????????????????????????????????????????????????????????????
CONCLUSION
The use of SiRNAs as therapeutic agents has changed the landscape of drug development. An extensive assessment of the distinct clinical pharmacology, safety, and efficacy characteristics of this new class of medicines is being actively sought after. The FDA-approved siRNA treatments have distinct clinical pharmacology features when compared to small compounds and protein-based medicines, as this study explains. Due to miRNA's special ability to target numerous mRNA transcripts, a single medication can now regulate several targets or biochemical pathways?many of which are dysregulated in disease. To do this, thorough basic and clinical research is required to address aspects including dose, cross-reactivity, on-target efficacy, and undesired effects before they can receive regulatory approval. Numerous RNA therapies have been developed to treat a variety of ailments, and their approval for use in treating a range of diseases is a result of the encouraging outcomes of numerous clinical trials. But the main obstacle to the development of RNA therapies for many diseases is the absence of appropriate delivery mechanisms.
REFERENCE



Sakshi Bodke, Mayuri Jagtap*, Avinash Darekar, RNA Therapeutics: Clinical Pharmacokinetic and Therapeutic Monitoring, Int. J. Sci. R. Tech., 2024, 1(3) 25-30. https://doi.org/10.5281/zenodo.13884776
 10.5281/zenodo.13884776
10.5281/zenodo.13884776